Mechanism of Action
From In-Silico Design to Structural Validation.
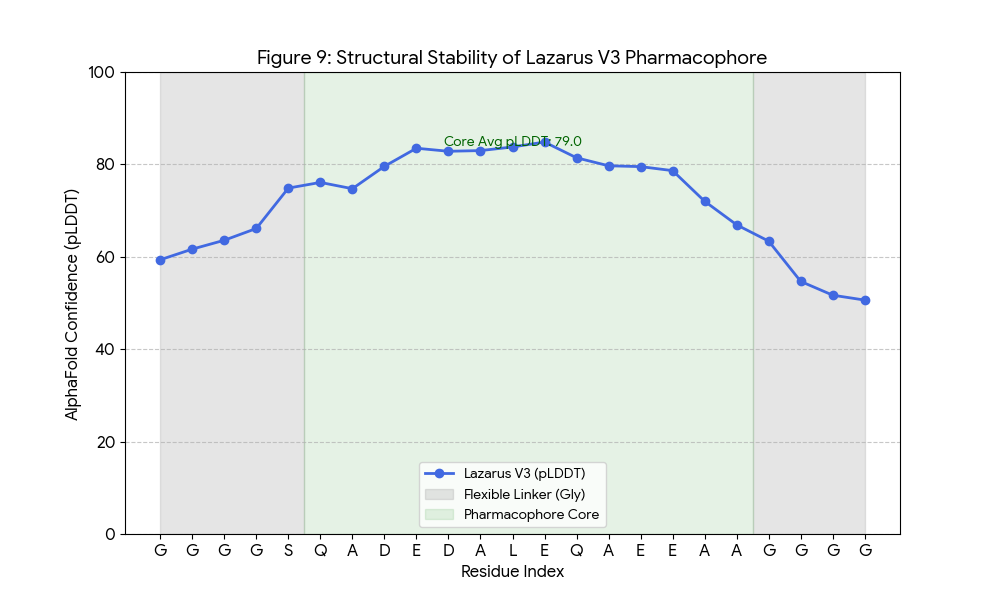
Designed for Rigidity
Lazarus-V3 was engineered using AlphaFold 3 to form a stable alpha-helix only upon target binding. The peptide maintains high confidence scores in the bioactive core, ensuring precise docking.
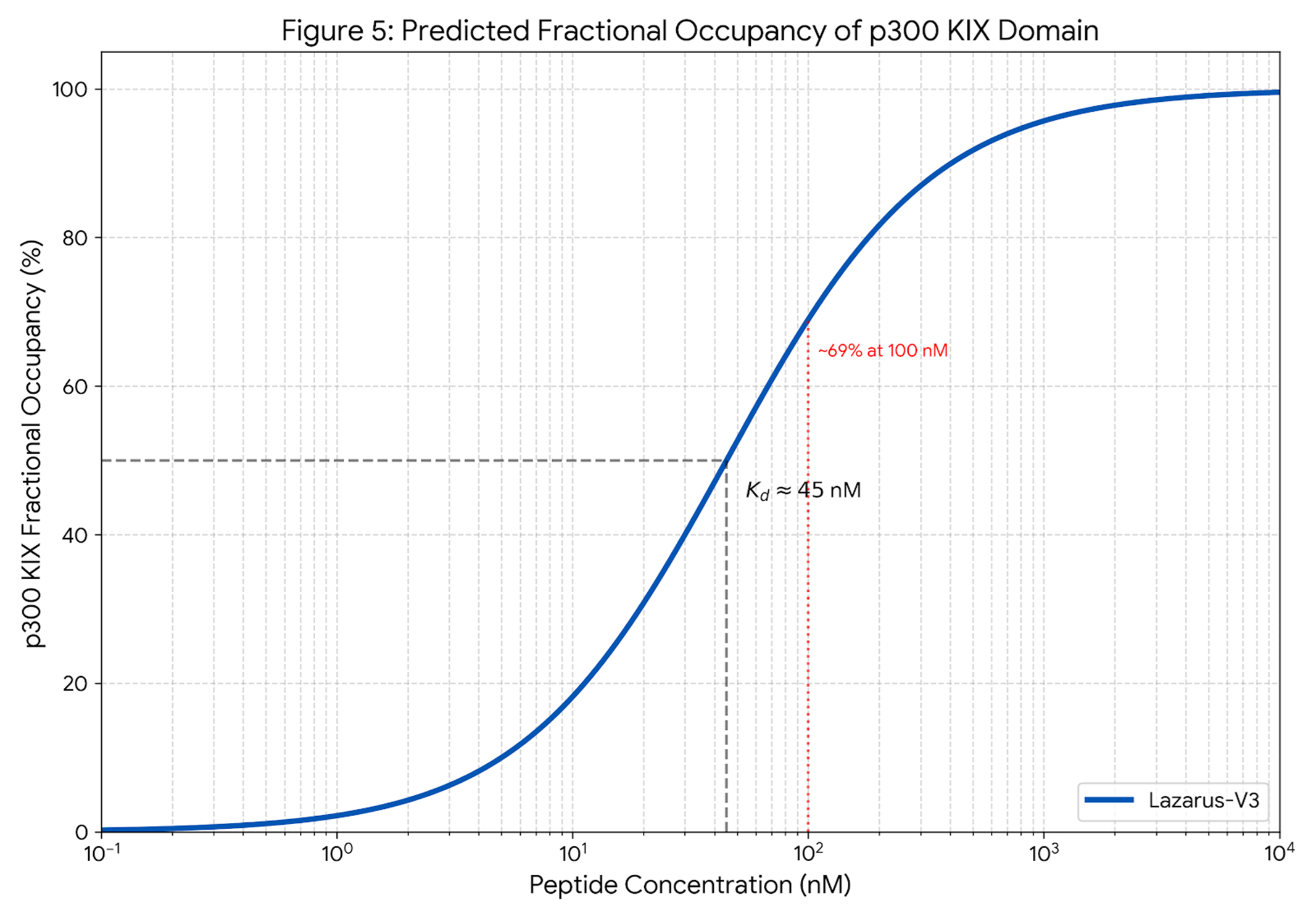
High-Affinity Capture
Dose-response modeling predicts a dissociation constant (Kd) in the nanomolar range. This affinity is tuned to outcompete natural ligands (like HIF-1α) only in the overexpression context of the tumor.
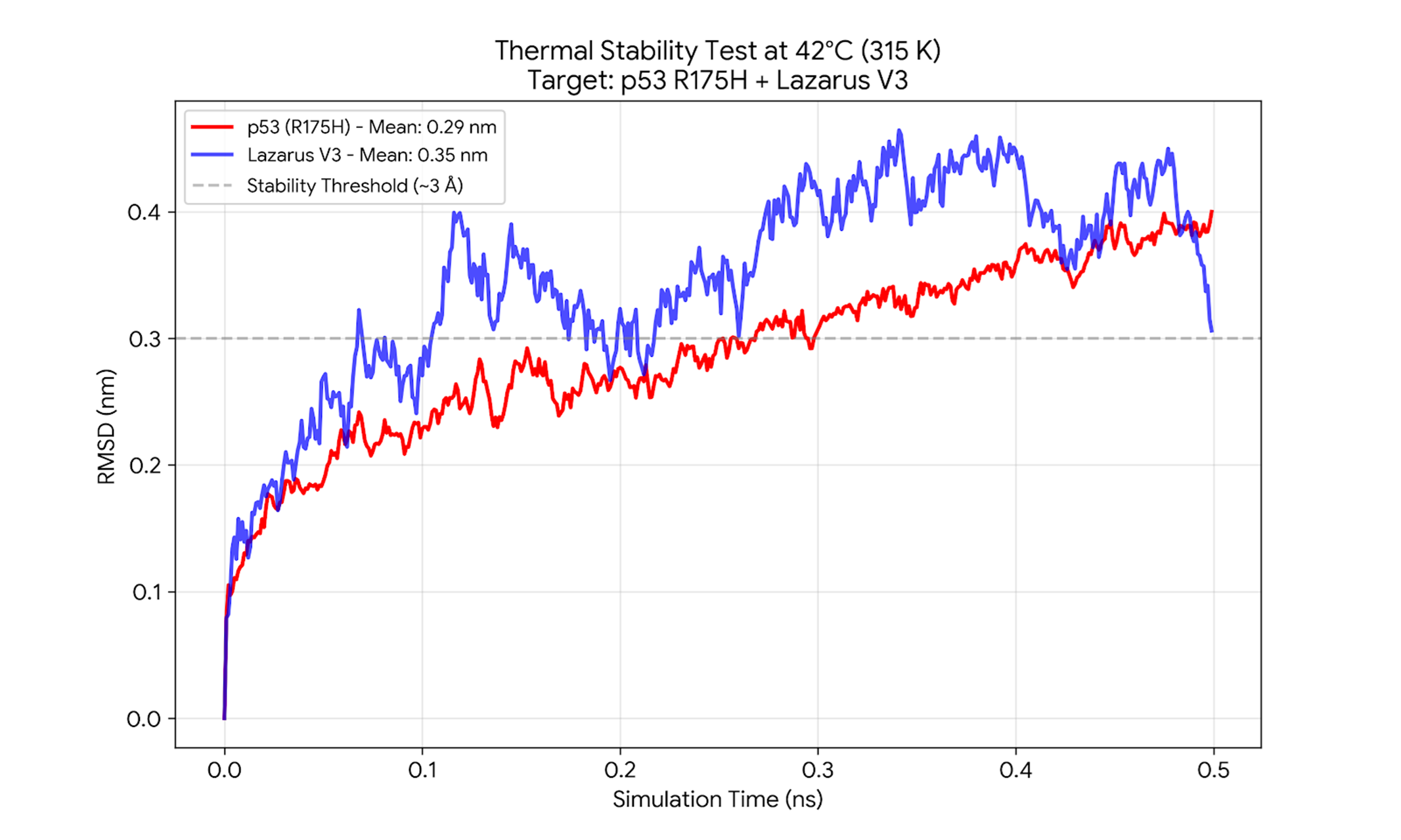
The "Solubility Chaperone" Effect
Molecular Dynamics simulations at 315K (42°C) confirm that Lazarus-V3 prevents the thermal unfolding of the p53-R175H mutant. The complex maintains a low RMSD, effectively "rescuing" the structure from aggregation.